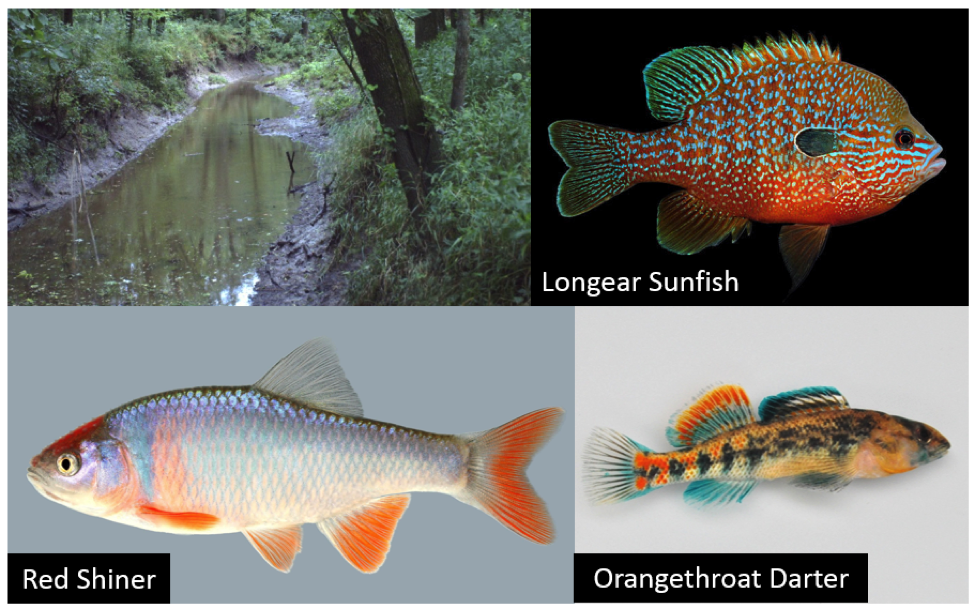
Stream fish and altered flow regimes in MO
By Nick Sievert, School of Natural Resources Nick participated in the event in Montgomery The small, muddy stream running through a woodlot on a central Missouri farm might not look like a hotbed of biodiversity, but chances are it has over a dozen different species of fish calling it home. Some like the Orangethroat Darter,…